Resources: Difference between revisions
imported>Ctelmer No edit summary |
No edit summary |
||
| (3 intermediate revisions by 2 users not shown) | |||
| Line 7: | Line 7: | ||
Please read following post to learn about our BACs resource and experimental methods<br /> | Please read following post to learn about our BACs resource and experimental methods<br /> | ||
[[recombac|Recombinant Bacs]]<br /> | [[recombac|Recombinant Bacs]]<br /> | ||
Annotation of BAC Sequences | Annotation of BAC Sequences with Sea Urchin Gene AnnotatoR (SUGAR, no longer available) | ||
| Line 13: | Line 13: | ||
A virtual map of the genome was constructed by sequencing the ends of 76,020 BAC recombinants (average length 125kb). The BAC-end sequence tag connectors (STCs) occur an average of 10kb apart. They can be used to assemble contigs surrounding any gene of interest. Using Blast matches to sequences from Bac-ends and complete BACs, confirmations from cDNA sequences we estimate that the sea urchin genome contains a total of 22+/-5X10<sup>3</sup> genes. | A virtual map of the genome was constructed by sequencing the ends of 76,020 BAC recombinants (average length 125kb). The BAC-end sequence tag connectors (STCs) occur an average of 10kb apart. They can be used to assemble contigs surrounding any gene of interest. Using Blast matches to sequences from Bac-ends and complete BACs, confirmations from cDNA sequences we estimate that the sea urchin genome contains a total of 22+/-5X10<sup>3</sup> genes. | ||
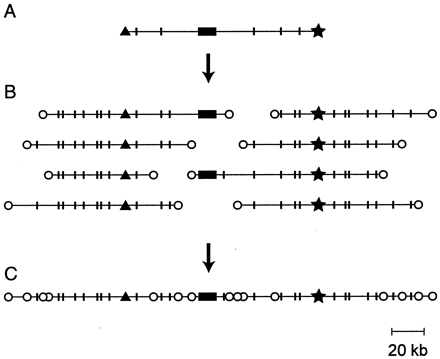
[[ | [[File:VirtualMap.gif|frame|left|Recovery of a contig surrounding a known gene by use of the STC database. (A) The arrayed BAC library is screened by using a cDNA probe, and a BAC containing the gene sequence is selected; the most desirable clone would be the longest that has the gene sequence (solid box) toward the middle. The STCs for that BAC (black-triangle, star) are recovered from the database together with the length distribution of restriction fragments produced by the sites (indicated by short, vertical lines). (B) Oligonucleotide probes are created from the left and right STCs and used to scan the BAC library. The selected clones are aligned to overlap in the restriction digest patterns. The probes should not lie within repetitive sequences: if they do, too many BACs will score positive, and their restriction fragment lengths will not align. (C) The clones extending the longest distance left and right plus the original clone (A) constitute a contig surrounding the gene, marked by STC tags at frequent intervals.]]. | ||
'''Repeats'''<br /> | '''Repeats'''<br /> | ||
| Line 28: | Line 28: | ||
'''cDNA sequences'''<br /> | '''cDNA sequences'''<br /> | ||
Approximately 13,000 cDNA sequences were obtained from the primary mesenchyme cell library. These sequences comprise 7,400 unique sequences when all of the overlaps are assembled. When these are searched against the BAC-end sequences, 1087 unique matches occur. Thus, the sequence matches between the BAC-ends, the ESTs, and the published data bases all give results commensurate with the conclusion that the collection of sequences we have obtained are of a quality suitable for gene discovery investigations in the sea urchin embryo. <span class="newwin">[http:// | Approximately 13,000 cDNA sequences were obtained from the primary mesenchyme cell library. These sequences comprise 7,400 unique sequences when all of the overlaps are assembled. When these are searched against the BAC-end sequences, 1087 unique matches occur. Thus, the sequence matches between the BAC-ends, the ESTs, and the published data bases all give results commensurate with the conclusion that the collection of sequences we have obtained are of a quality suitable for gene discovery investigations in the sea urchin embryo. <span class="newwin">[http://www.echinobase.org/literature/article.do?method=display&articleId=37498 (Cameron et al., Proc. Natl. Acad. Sci. USA, Vol. 97, Issue 17, 9514-9518, August 15, 2000 )]</span> | ||
| Line 48: | Line 48: | ||
'''Sea Urchin Codon Usage Table'''<br /> | '''Sea Urchin Codon Usage Table'''<br /> | ||
The Sea Urchin codon usage table can be accessed [http://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=7668 here] | The Sea Urchin codon usage table can be accessed [http://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=7668 here]. | ||
'''The principal investigators who participated in establishment of the Sea Urchin Genome Project:'''<br /> | |||
<p>Dr. Eric Davidson, California Institute of Technology</p> | |||
<p>Dr. Charles Ettensohn, Carnegie-Mellon University</p> | |||
<p>Dr. Leroy Hood, Institute for Systems Biology, Seattle Washington</p> | |||
<p>Dr. Greg Wray Duke University, The Sea Urchin Genome Project</p> | |||
<p>Dr. Andrew Cameron, California Institute of Technology</p> | |||
'''Advisory Committee for the HGRI Sea Urchin Sequencing Project:'''<br /> | |||
<p>Eric H. Davidson, California Institute of Technology</p> | |||
<p>R. Andrew Cameron, California Institute of Technology</p> | |||
<p>Robert C. Angerer, National Institute of Dental and Craniofacial Research</p> | |||
<p>Lynne Angerer, National Institute of Dental and Craniofacial Research</p> | |||
<p>James A. Coffman, Mount Desert Island Biological Laboratory</p> | |||
<p>William H. Klein, M. D. Anderson Cancer Center</p> | |||
<p>Donal Manahan, University of Southern California</p> | |||
<p>David R. McClay, Duke University</p> | |||
<p>Jonathan P. Rast, University of Toronto</p> | |||
<p>Victor D. Vacquier, Scripps Institute of Oceanography</p> | |||
Latest revision as of 15:03, 6 October 2023
The combined resources provided by the preliminary Sea Urchin Genome Project are included on this site. The individual resources listed below are useful for gene discovery approaches, expressed sequence tag analysis and most importantly, studies of gene regulation in the sea urchin.
BAC Table
BAC Table can be accessed here BAC Table
Please read following post to learn about our BACs resource and experimental methods
Recombinant Bacs
Annotation of BAC Sequences with Sea Urchin Gene AnnotatoR (SUGAR, no longer available)
BACs, BAC-ends and Gene Number
A virtual map of the genome was constructed by sequencing the ends of 76,020 BAC recombinants (average length 125kb). The BAC-end sequence tag connectors (STCs) occur an average of 10kb apart. They can be used to assemble contigs surrounding any gene of interest. Using Blast matches to sequences from Bac-ends and complete BACs, confirmations from cDNA sequences we estimate that the sea urchin genome contains a total of 22+/-5X103 genes.
.
Repeats
Since the first sea urchin genomic sequencing project was undertaken, a number of collections of the repeat sequences in the purple sea urchin have been collected. A survey of the 76,000 BAC ends for repeat sequences is described here.
cDNA Libraries
We maintain a suite of cDNA libraries from embryonic stages, larval stages and adult tissues of the reference species, Strongylocentrotus purpuratus. A few cDNA libraries from other species are also available (see Table) These libraries are stored in 384-well plates permitting easy replication and re-spotting as needed. The libraries are spotted onto filters for screening. Library filters are available to members of the research community. (How to order). These libraries have been used as the basis for EST sequencing as part of the original genome sequencing project reported in Science. Several other smaller EST projects have also been completed (for example: Zhu, X., Mahairas, G., Cameron, R. A., Davidson, E. H. and Ettensohn, C. A. Large-scale analysis of mRNAs expressed by primary mesenchyme cells of the sea urchin embryo. Development 128, 2615-2627, 2001). The clones sequenced in these projects are accessioned into Genbank and SpBase with the plate and well locations attached. Thus one can use sequence search programs to find the clones and then request them from our Resource. In other words "cloning by computer" is enabled here.
BAC Libraries
Genomic sequence segments are maintained in bacterial artificial chromosome (BAC) libraries. We currently have libraries for seven species of lower deuterostomes in our resource (see Table)The vector used is pBACe3.6 which originates from Children's Hospital Oakland Research Institute in Oakland, California, USA. This vector has chloramphenicol antibiotic resistance and is fully described in an article by Frengen and colleages. The vector sequence is available here. The average insert size for our BAC libraries is about 140 Kb thus 1X coverage of the 800 Megabase genome of S. purpuratus is on average 5700 clones. Our libraries are at least 100,000 clones providing about 17X genome coverage.
cDNA sequences
Approximately 13,000 cDNA sequences were obtained from the primary mesenchyme cell library. These sequences comprise 7,400 unique sequences when all of the overlaps are assembled. When these are searched against the BAC-end sequences, 1087 unique matches occur. Thus, the sequence matches between the BAC-ends, the ESTs, and the published data bases all give results commensurate with the conclusion that the collection of sequences we have obtained are of a quality suitable for gene discovery investigations in the sea urchin embryo. (Cameron et al., Proc. Natl. Acad. Sci. USA, Vol. 97, Issue 17, 9514-9518, August 15, 2000 )
Gene Model
During sea urchin genome project, several groups came up with gene predictions based on diverse approaches (ab initio, homology-based or empirical). Baylor used GLEAN methodology to combine those gene-sets into 28,944 unique genes. Their structures were derived from V0.5 genome assembly. At SpBase, we adopted Baylor's GLEAN genes and renamed each GLEAN IDs as GLEAN3_12345 to SPU_012345. New SPU genes will be added with IDs starting from 030000. This first release only modified the gene IDs from GLEAN and adopted them into SpBase. No real changes of gene structures were done.
Genome sequence traces
Beginning in March 2003, the Baylor College of Medicine, Human Genome Sequencing center (www.hgsc.bcm.tmc.edu) began to produce sea urchin sequences. First, a whole genome shotgun (WGS) project was undertaken and the individual sequences are deposited in the Genbank Trace Repository at NCBI. We have downloaded these traces,analyzed them by Blast and posted the matches in a searchable form on this web site. We will continue to do so until assembled genome sequences are posted at NCBI.
Quantitative PCR primers
The Davidson laboratory at Caltech has generated a panel of quantitative PCR primers useful for measuring the level of mRNA abundance for genes involved in early development in general and the endomesoderm gene regulatory network in particular. A table of primer sequences and comments can be viewed (here).
Nanostring Codeset
The Nanostring nCounter identifies and counts RNA molecules based on a fluorescent barcode attached to a sequence specific hybridization (Geiss, G. K. et al,2008). Our newly designed probe set contains codes for 341 genes covering the majority of active, and spatially restricted regulatory genes in the Strongylocentrotus purpuratus embryo up to pluteus (72 h post-fertilization). A description of the use of our previous code set was published by (Materna et al.). They showed that Nanostring nCounter yields measurements with high fidelity over 5 orders of magnitude at levels down to a few transcripts per embryo. The genes and sequences in the new codeset are tabulated here.
Sea Urchin Codon Usage Table
The Sea Urchin codon usage table can be accessed here.
The principal investigators who participated in establishment of the Sea Urchin Genome Project:
Dr. Eric Davidson, California Institute of Technology
Dr. Charles Ettensohn, Carnegie-Mellon University
Dr. Leroy Hood, Institute for Systems Biology, Seattle Washington
Dr. Greg Wray Duke University, The Sea Urchin Genome Project
Dr. Andrew Cameron, California Institute of Technology
Advisory Committee for the HGRI Sea Urchin Sequencing Project:
Eric H. Davidson, California Institute of Technology
R. Andrew Cameron, California Institute of Technology
Robert C. Angerer, National Institute of Dental and Craniofacial Research
Lynne Angerer, National Institute of Dental and Craniofacial Research
James A. Coffman, Mount Desert Island Biological Laboratory
William H. Klein, M. D. Anderson Cancer Center
Donal Manahan, University of Southern California
David R. McClay, Duke University
Jonathan P. Rast, University of Toronto
Victor D. Vacquier, Scripps Institute of Oceanography